Neural-network based simulation of rare event processes at the water/oxide interface
Subproject P12
Atomistic computer simulations of processes occurring at the water/oxide interface are challenging in several ways. The calculation of atomic forces based on ab initio methods is computationally very demanding, and barrier crossing events may lead to long computation times. Both these aspects severely limit accessible system sizes and simulation times.
In project P12, we will address these challenges using a combination of machine learning and advanced rare event sampling methods. In particular, using software developed in our group and collaborating with P03 Kresse, we will train neural network potentials based on the Behler-Parrinello approach for oxide/water interfaces, starting with the Fe3O4/water system studied in P11 Backus. We will pay special attention to error estimation and the correct treatment of long-range interactions. With the new potential, we will study the structure and dynamics of water near the oxide surface to provide the atomistic information necessary to rationalize the spectroscopy experiments of P11 Backus. Another important goal of P12 is to explore how deep generative models can be used to enhance rare event simulations. For this purpose, we will apply normalizing flows, represented by deep neural networks, to trajectory space. The resulting improved transition path sampling simulations will be used to study reactive processes investigated experimentally in other subprojects of TACO.
Expertise
Our research efforts focus on the development of simulation algorithms and their application to investigate dynamical processes in condensed matter systems based on the principles of equilibrium and non-equilibrium statistical mechanics. In particular, we have helped to create the transition path sampling methodology for the simulation of rare but important events, such as nucleation aprocesses, chemical reactions and biomolecular reorganizations. More recently, we have worked on applying machine learning methods to molecular structure recognition and the representation of potential and free energy surfaces.
Recent research topics include:
- Self-assembly of nanocrystals
- Folding and unfolding of biopolymers
- Interfaces in aqueous systems
- Phase separation in alloys
- Thermo-polarisation
- Structure and dynamics of water and ice
- Cavitation
- Crystallization
- Non-equilibrium work fluctuations
Team
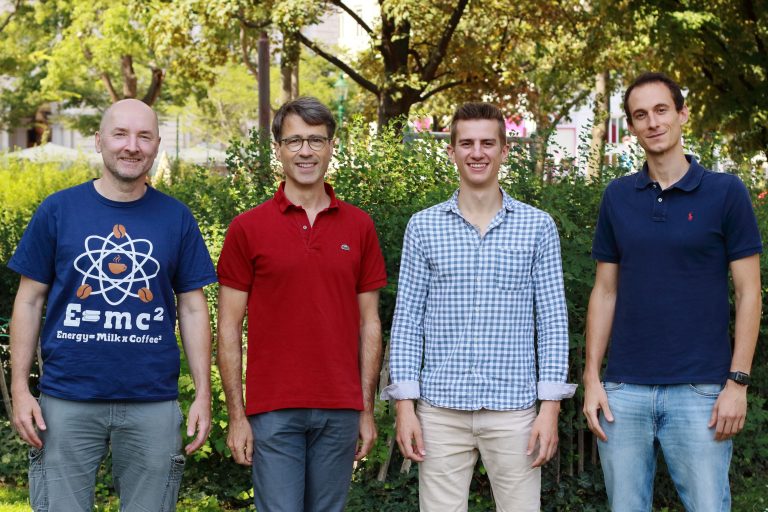

Associates
Publications
2024
de Hijes, Pablo Montero; Dellago, Christoph; Jinnouchi, Ryosuke; Schmiedmayer, Bernhard; Kresse, Georg
Journal ArticleOpen AccessIn: The Journal of Chemical Physics, vol. 160, iss. 11, no. 114107, 2024.
Abstract | Links | BibTeX | Tags: P03, P12
@article{10.1063/5.0197105,
title = {Comparing machine learning potentials for water: Kernel-based regression and Behler–Parrinello neural networks},
author = {Pablo Montero de Hijes and Christoph Dellago and Ryosuke Jinnouchi and Bernhard Schmiedmayer and Georg Kresse},
doi = {https://doi.org/10.1063/5.0197105},
year = {2024},
date = {2024-03-20},
urldate = {2024-03-20},
journal = {The Journal of Chemical Physics},
volume = {160},
number = {114107},
issue = {11},
abstract = {In this paper, we investigate the performance of different machine learning potentials (MLPs) in predicting key thermodynamic properties of water using RPBE + D3. Specifically, we scrutinize kernel-based regression and high-dimensional neural networks trained on a highly accurate dataset consisting of about 1500 structures, as well as a smaller dataset, about half the size, obtained using only on-the-fly learning. This study reveals that despite minor differences between the MLPs, their agreement on observables such as the diffusion constant and pair-correlation functions is excellent, especially for the large training dataset. Variations in the predicted density isobars, albeit somewhat larger, are also acceptable, particularly given the errors inherent to approximate density functional theory. Overall, this study emphasizes the relevance of the database over the fitting method. Finally, this study underscores the limitations of root mean square errors and the need for comprehensive testing, advocating the use of multiple MLPs for enhanced certainty, particularly when simulating complex thermodynamic properties that may not be fully captured by simpler tests.},
keywords = {P03, P12},
pubstate = {published},
tppubtype = {article}
}
Falkner, Sebastian; Coretti, Alessandro; Dellago, Christoph
Journal ArticleOpen AccessIn: Physical Review Letters, vol. 132, iss. 12, pp. 128001, 2024.
Abstract | Links | BibTeX | Tags: P12
@article{Falkner2024,
title = {Enhanced Sampling of Configuration and Path Space in a Generalized Ensemble by Shooting Point Exchange},
author = {Sebastian Falkner and Alessandro Coretti and Christoph Dellago},
url = {https://arxiv.org/abs/2302.08757},
doi = {https://doi.org/10.1103/PhysRevLett.132.128001},
year = {2024},
date = {2024-03-18},
urldate = {2024-03-18},
journal = {Physical Review Letters},
volume = {132},
issue = {12},
pages = {128001},
abstract = {The computer simulation of many molecular processes is complicated by long timescales caused by rare transitions between long-lived states. Here, we propose a new approach to simulate such rare events, which combines transition path sampling with enhanced exploration of configuration space. The method relies on exchange moves between configuration and trajectory space, carried out based on a generalized ensemble. This scheme substantially enhances the efficiency of the transition path sampling simulations, particularly for systems with multiple transition channels, and yields information on thermodynamics, kinetics and reaction coordinates of molecular processes without distorting their dynamics. The method is illustrated using the isomerization of proline in the KPTP tetrapeptide.},
keywords = {P12},
pubstate = {published},
tppubtype = {article}
}
Gorfer, Alexander; Abart, Rainer; Dellago, Christoph
Structure and thermodynamics of defects in Na-feldspar from a neural network potential
Journal ArticleSubmittedarXivIn: arXiv, 2024.
Abstract | Links | BibTeX | Tags: P12
@article{Gorfer_2024,
title = {Structure and thermodynamics of defects in Na-feldspar from a neural network potential},
author = {Alexander Gorfer and Rainer Abart and Christoph Dellago},
url = {https://arxiv.org/abs/2402.14640},
year = {2024},
date = {2024-02-22},
urldate = {2024-02-22},
journal = {arXiv},
abstract = {The diffusive phase transformations occurring in feldspar, a common mineral in the crust of the Earth, are essential for reconstructing the thermal histories of magmatic and metamorphic rocks. Due to the long timescales over which these transformations proceed, the mechanism responsible for sodium diffusion and its possible anisotropy has remained a topic of debate. To elucidate this defect-controlled process, we have developed a Neural Network Potential (NNP) trained on first-principle calculations of Na-feldspar (Albite) and its charged defects. This new force field reproduces various experimentally known properties of feldspar, including its lattice parameters, elastic constants as well as heat capacity and DFT-calculated defect formation energies. A new type of dumbbell interstitial defect is found to be most favorable and its free energy of formation at finite temperature is calculated using thermodynamic integration. The necessity of including electrostatic corrections before training an NNP is demonstrated by predicting more consistent defect formation energies.},
keywords = {P12},
pubstate = {published},
tppubtype = {article}
}

Omranpour, Amir; de Hijes, Pablo Montero; Behler, Jörg; Dellago, Christoph
Perspective: Atomistic Simulations of Water and Aqueous Systems with Machine Learning Potentials
Journal ArticleSubmittedarXivIn: arXiv, 2024.
Abstract | Links | BibTeX | Tags: P12
@article{Omranpour_2024,
title = {Perspective: Atomistic Simulations of Water and Aqueous Systems with Machine Learning Potentials},
author = {Amir Omranpour and Pablo Montero de Hijes and Jörg Behler and Christoph Dellago},
url = {https://arxiv.org/abs/2401.17875},
year = {2024},
date = {2024-01-31},
urldate = {2024-01-31},
journal = {arXiv},
abstract = {As the most important solvent, water has been at the center of interest since the advent of computer simulations. While early molecular dynamics and Monte Carlo simulations had to make use of simple model potentials to describe the atomic interactions, accurate ab initio molecular dynamics simulations relying on the first-principles calculation of the energies and forces have opened the way to predictive simulations of aqueous systems. Still, these simulations are very demanding, which prevents the study of complex systems and their properties. Modern machine learning potentials (MLPs) have now reached a mature state, allowing to overcome these limitations by combining the high accuracy of electronic structure calculations with the efficiency of empirical force fields. In this Perspective we give a concise overview about the progress made in the simulation of water and aqueous systems employing MLPs, starting from early work on free molecules and clusters via bulk liquid water to electrolyte solutions and solid-liquid interfaces.},
keywords = {P12},
pubstate = {published},
tppubtype = {article}
}
2023

Domenichini, Giorgio; Dellago, Christoph
Molecular Hessian matrices from a machine learning random forest regression algorithm
Journal ArticleOpen AccessIn: The Journal of Chemical Physics, vol. 159, iss. 19, no. 194111, 2023.
Abstract | Links | BibTeX | Tags: P12
@article{10.1063/5.0169384,
title = {Molecular Hessian matrices from a machine learning random forest regression algorithm},
author = {Giorgio Domenichini and Christoph Dellago},
doi = {https://doi.org/10.1063/5.0169384},
year = {2023},
date = {2023-11-20},
urldate = {2023-11-20},
journal = {The Journal of Chemical Physics},
volume = {159},
number = {194111},
issue = {19},
abstract = {In this article, we present a machine learning model to obtain fast and accurate estimates of the molecular Hessian matrix. In this model, based on a random forest, the second derivatives of the energy with respect to redundant internal coordinates are learned individually. The internal coordinates together with their specific representation guarantee rotational and translational invariance. The model is trained on a subset of the QM7 dataset but is shown to be applicable to larger molecules picked from the QM9 dataset. From the predicted Hessian, it is also possible to obtain reasonable estimates of the vibrational frequencies, normal modes, and zero point energies of the molecules.},
keywords = {P12},
pubstate = {published},
tppubtype = {article}
}

Falkner, Sebastian; Coretti, Alessandro; Romano, Salvatore; Geissler, Phillip L.; Dellago, Christoph
Conditioning Boltzmann generators for rare event sampling
Journal ArticleOpen AccessIn: Machine Learning: Science and Technology, vol. 4, iss. 3, no. 035050, 2023.
Abstract | Links | BibTeX | Tags: P12
@article{Falkner_2023,
title = {Conditioning Boltzmann generators for rare event sampling},
author = {Sebastian Falkner and Alessandro Coretti and Salvatore Romano and Phillip L. Geissler and Christoph Dellago},
url = {https://arxiv.org/abs/2207.14530},
doi = {10.1088/2632-2153/acf55c},
year = {2023},
date = {2023-09-22},
urldate = {2023-09-22},
journal = {Machine Learning: Science and Technology},
volume = {4},
number = {035050},
issue = {3},
abstract = {Understanding the dynamics of complex molecular processes is often linked to the study of infrequent transitions between long-lived stable states. The standard approach to the sampling of such rare events is to generate an ensemble of transition paths using a random walk in trajectory space. This, however, comes with the drawback of strong correlations between subsequently sampled paths and with an intrinsic difficulty in parallelizing the sampling process. We propose a transition path sampling scheme based on neural-network generated configurations. These are obtained employing normalizing flows, a neural network class able to generate statistically independent samples from a given distribution. With this approach, not only are correlations between visited paths removed, but the sampling process becomes easily parallelizable. Moreover, by conditioning the normalizing flow, the sampling of configurations can be steered towards regions of interest. We show that this approach enables the resolution of both the thermodynamics and kinetics of the transition region for systems that can be sampled using exact-likelihood generative models.},
keywords = {P12},
pubstate = {published},
tppubtype = {article}
}

Hijes, Pablo Montero; Romano, Salvatore; Gorfer, Alexander; Dellago, Christoph
The kinetics of the ice–water interface from ab initio machine learning simulations
Journal ArticleOpen AccessIn: The Journal of Chemical Physics, vol. 158, no. 204706, 2023.
Abstract | Links | BibTeX | Tags: P12
@article{Hijes2023a,
title = {The kinetics of the ice–water interface from \textit{ab initio} machine learning simulations},
author = {Pablo Montero Hijes and Salvatore Romano and Alexander Gorfer and Christoph Dellago},
doi = {10.1063/5.0151011},
year = {2023},
date = {2023-05-24},
urldate = {2023-05-24},
journal = {The Journal of Chemical Physics},
volume = {158},
number = {204706},
publisher = {AIP Publishing},
abstract = {Molecular simulations employing empirical force fields have provided valuable knowledge about the ice growth process in the past decade. The development of novel computational techniques allows us to study this process, which requires long simulations of relatively large systems, with ab initio accuracy. In this work, we use a neural-network potential for water trained on the revised Perdew–Burke–Ernzerhof functional to describe the kinetics of the ice–water interface. We study both ice melting and growth processes. Our results for the ice growth rate are in reasonable agreement with previous experiments and simulations. We find that the kinetics of ice melting presents a different behavior (monotonic) than that of ice growth (non-monotonic). In particular, a maximum ice growth rate of 6.5 Å/ns is found at 14 K of supercooling. The effect of the surface structure is explored by investigating the basal and primary and secondary prismatic facets. We use the Wilson–Frenkel relation to explain these results in terms of the mobility of molecules and the thermodynamic driving force. Moreover, we study the effect of pressure by complementing the standard isobar with simulations at a negative pressure (−1000 bar) and at a high pressure (2000 bar). We find that prismatic facets grow faster than the basal one and that pressure does not play an important role when the speed of the interface is considered as a function of the difference between the melting temperature and the actual one, i.e., to the degree of either supercooling or overheating.},
keywords = {P12},
pubstate = {published},
tppubtype = {article}
}

Jung, Hendrik; Covino, Roberto; Arjun, A.; Leitold, Christian; Dellago, Christoph; Bolhuis, Peter G.; Hummer, Gerhard
Machine-guided path sampling to discover mechanisms of molecular self-organization
Journal ArticleOpen AccessIn: Nature Computational Science, vol. 3, pp. 334–345, 2023.
Abstract | Links | BibTeX | Tags: P12
@article{Jung2023,
title = {Machine-guided path sampling to discover mechanisms of molecular self-organization},
author = {Hendrik Jung and Roberto Covino and A. Arjun and Christian Leitold and Christoph Dellago and Peter G. Bolhuis and Gerhard Hummer},
doi = {10.1038/s43588-023-00428-z},
year = {2023},
date = {2023-04-24},
urldate = {2023-04-24},
journal = {Nature Computational Science},
volume = {3},
pages = {334--345},
publisher = {Springer Science and Business Media LLC},
abstract = {Molecular self-organization driven by concerted many-body interactions produces the ordered structures that define both inanimate and living matter. Here we present an autonomous path sampling algorithm that integrates deep learning and transition path theory to discover the mechanism of molecular self-organization phenomena. The algorithm uses the outcome of newly initiated trajectories to construct, validate and—if needed—update quantitative mechanistic models. Closing the learning cycle, the models guide the sampling to enhance the sampling of rare assembly events. Symbolic regression condenses the learned mechanism into a human-interpretable form in terms of relevant physical observables. Applied to ion association in solution, gas-hydrate crystal formation, polymer folding and membrane-protein assembly, we capture the many-body solvent motions governing the assembly process, identify the variables of classical nucleation theory, uncover the folding mechanism at different levels of resolution and reveal competing assembly pathways. The mechanistic descriptions are transferable across thermodynamic states and chemical space.},
keywords = {P12},
pubstate = {published},
tppubtype = {article}
}

Hijes, Pablo Montero; Espinosa, Jorge R; Vega, Carlos; Dellago, Christoph
Minimum in the pressure dependence of the interfacial free energy between ice Ih and water
Journal ArticleOpen AccessIn: The Journal of Chemical Physics, vol. 158, no. 12, pp. 124503, 2023.
Abstract | Links | BibTeX | Tags: P12
@article{Hijes2023,
title = {Minimum in the pressure dependence of the interfacial free energy between ice Ih and water},
author = {Pablo Montero Hijes and Jorge R Espinosa and Carlos Vega and Christoph Dellago},
doi = {10.1063/5.0140814},
year = {2023},
date = {2023-03-23},
urldate = {2023-03-23},
journal = {The Journal of Chemical Physics},
volume = {158},
number = {12},
pages = {124503},
publisher = {AIP Publishing},
abstract = {Despite the importance of ice nucleation, this process has been barely explored at negative pressures. Here, we study homogeneous ice nucleation in stretched water by means of molecular dynamics seeding simulations using the TIP4P/Ice model. We observe that the critical nucleus size, interfacial free energy, free energy barrier, and nucleation rate barely change between isobars from −2600 to 500 bars when they are represented as a function of supercooling. This allows us to identify universal empirical expressions for homogeneous ice nucleation in the pressure range from −2600 to 500 bars. We show that this universal behavior arises from the pressure dependence of the interfacial free energy, which we compute by means of the mold integration technique, finding a shallow minimum around −2000 bars. Likewise, we show that the change in the interfacial free energy with pressure is proportional to the excess entropy and the slope of the melting line, exhibiting in the latter a reentrant behavior also at the same negative pressure. Finally, we estimate the excess internal energy and the excess entropy of the ice Ih–water interface.},
keywords = {P12},
pubstate = {published},
tppubtype = {article}
}
2022
Tröster, Andreas; Verdi, Carla; Dellago, Christoph; Rychetsky, Ivan; Kresse, Georg; Schranz, Wilfried
Hard antiphase domain boundaries in strontium titanate unravelled using machine-learned force fields
Journal ArticleIn: Physical Review Materials, vol. 6, no. 9, pp. 094408, 2022.
Abstract | Links | BibTeX | Tags: P03, P12
@article{Troester2022,
title = {Hard antiphase domain boundaries in strontium titanate unravelled using machine-learned force fields},
author = {Andreas Tröster and Carla Verdi and Christoph Dellago and Ivan Rychetsky and Georg Kresse and Wilfried Schranz},
doi = {10.1103/physrevmaterials.6.094408},
year = {2022},
date = {2022-09-16},
urldate = {2022-09-16},
journal = {Physical Review Materials},
volume = {6},
number = {9},
pages = {094408},
publisher = {American Physical Society (APS)},
abstract = {We investigate the properties of hard antiphase boundaries in SrTiO_{3} using machine-learned force fields. In contrast to earlier findings based on standard \textit{ab initio} methods, for all pressures up to 120kbar the observed domain wall pattern maintains an almost perfect Néel character in quantitative agreement with Landau-Ginzburg-Devonshire theory, and the in-plane polarization P_{3} shows no tendency to decay to zero. Together with the switching properties of P_{3} under reversal of the Néel order parameter component, this provides hard evidence for the presence of rotopolar couplings. The present approach overcomes the severe limitations of \textit{ab initio} simulations of wide domain walls and opens avenues toward concise atomistic predictions of domain-wall properties even at finite temperatures.},
keywords = {P03, P12},
pubstate = {published},
tppubtype = {article}
}






